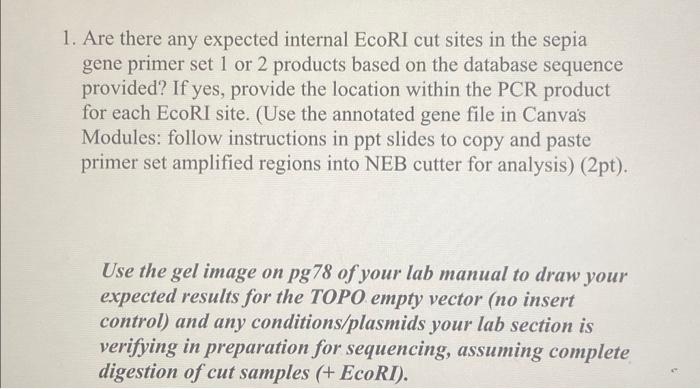
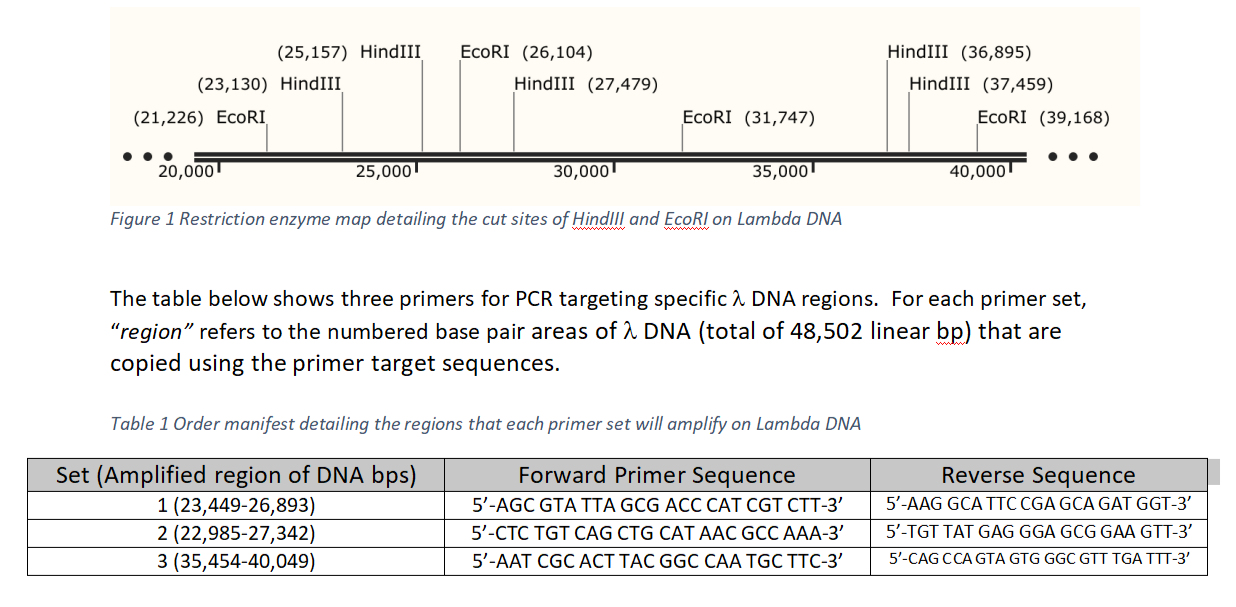
Schematic for adapter and primer design for the two rare cutters EcoRI and PstI and the frequent cutter MseI | Learn Science at Scitable

AFSM adapters and primers. (a, c) Sequences of three different types of... | Download Scientific Diagram
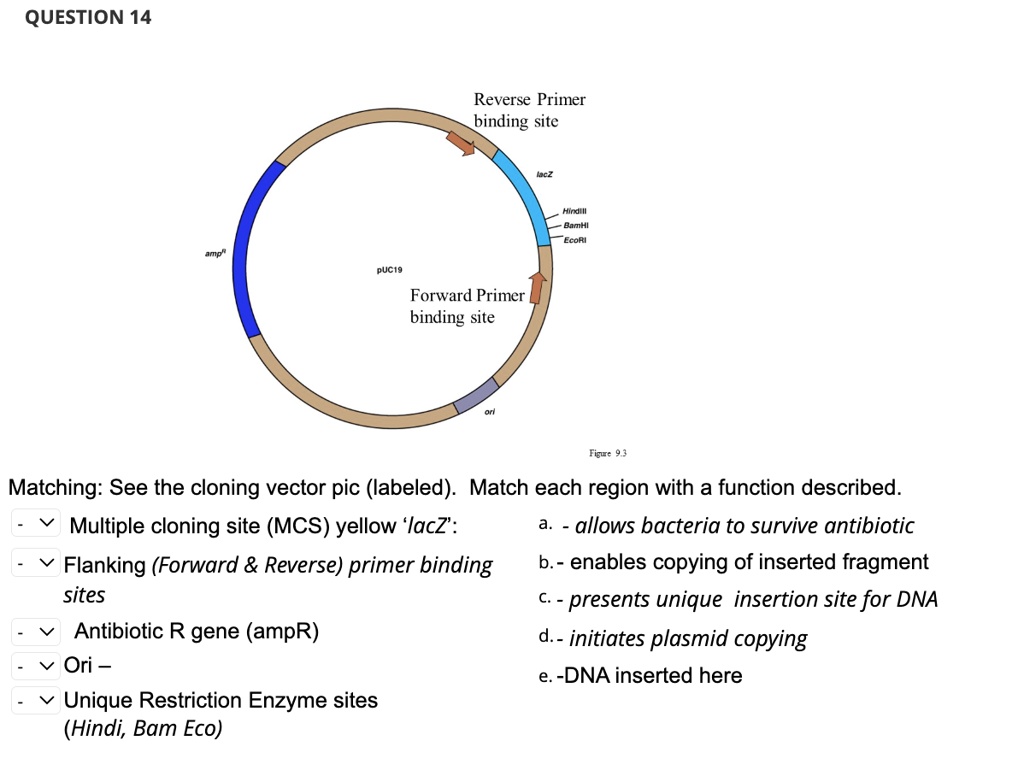
SOLVED: QUESTION 14 Reverse Primer binding site BamHI EcoRI pUC19 Forward Primer binding site Matching: See the cloning vector pic (labeled). Match each region with a function described. Multiple cloning site (MCS)
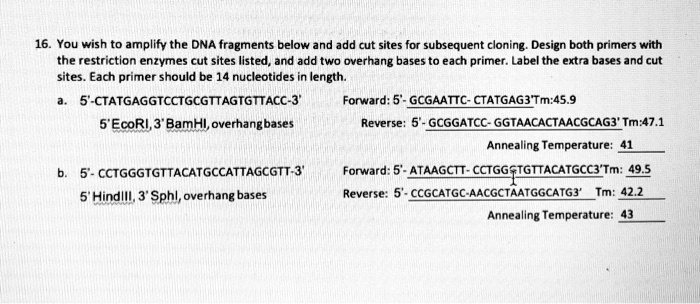

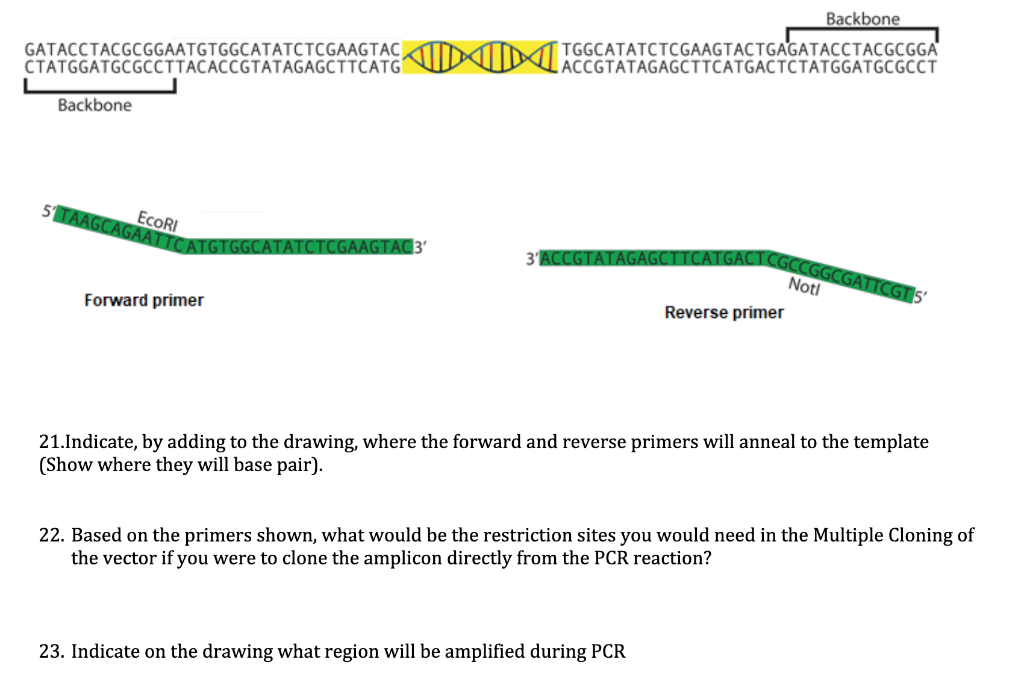
SOLVED: You wish to amplify the DNA fragments below and add cut sites for subsequent cloning. Design both primers with the restriction enzyme cut sites listed, and add two overhang bases to

The 373 bp DNA sequence between the EcoRI and RsaI sites from pUC8-31... | Download Scientific Diagram

a NcoI-Primer F DNA sequence and corresponding amino acids sequence... | Download Scientific Diagram